Nextflow Pipeline
For users interested in systematically comparing results across multiple moDiNA configurations, the Nextflow pipeline is the correct choice as it enables parallel, local execution using conda, Docker, Singularity, and distributed execution on HPC clusters using SLURM.
Installation
Note
If you are new to Nextflow, please refer to this page on how to set-up Nextflow.
Then you need to clone the moDiNA Nextflow pipeline repository:
git clone https://github.com/DyHealthNet/moDiNA_nf_pipeline.git
TODO: create conda environments if this is not working automatically…
Parameters
The Nextflow pipeline is designed to run multiple configurations in parallel, allowing users to systematically evaluate different settings and their impact on the results. To specify the configurations, you can either change the parameters in the nextflow.config file or create a configuration file (e.g., params.yml) that contains the parameters for each configuration you want to run.
Pipeline Parameters
This page documents all configurable parameters of the moDiNA Nextflow pipeline.
Parameters are set in nextflow.config or via a YAML params file (-params-file params.yml).
General Parameters
Parameter |
Type |
Default |
Description |
|---|---|---|---|
|
|
|
Label assigned to the first context (e.g. |
|
|
|
Label assigned to the second context (e.g. |
|
|
|
Determines how moDiNA configurations are specified. One of |
|
|
|
Source of input data. Either |
|
|
|
Absolute path to the directory where all pipeline outputs are written. |
|
|
|
Absolute path to the work directory of the Nextflow pipeline. |
|
|
|
Boolean specifying whether evaluation plots should be generated. |
Environment Parameters
Parameter |
Type |
Description |
|---|---|---|
|
|
Absolute path to the conda environment used for all core moDiNA processes (network inference, ranking, …). |
|
|
Absolute path to the conda environment used for R-based evaluation processes. |
Simulation Parameters (simulation.*)
Only required when data_type = 'simulation'.
Parameter |
Type |
Default |
Description |
|---|---|---|---|
|
|
|
Number of independent simulation replicates to run. |
|
|
|
Number of binary nodes to simulate. |
|
|
|
Number of continuous nodes to simulate. |
|
|
|
Number of categorical nodes to simulate. |
|
|
|
Number of samples (observations) per context. |
|
|
|
Number of continuous nodes with an artificially introduced mean shift between contexts. |
|
|
|
Number of binary nodes with an artificially introduced mean shift between contexts. |
|
|
|
Number of categorical nodes with an artificially introduced mean shift between contexts. |
|
|
|
Number of continuous–continuous node pairs with an artificially introduced correlation difference between contexts. |
|
|
|
Number of binary–binary node pairs with an artificially introduced correlation difference between contexts. |
|
|
|
Number of categorical–categorical node pairs with an artificially introduced correlation difference between contexts. |
|
|
|
Number of binary–continuous node pairs with an artificially introduced correlation difference between contexts. |
|
|
|
Number of binary–categorical node pairs with an artificially introduced correlation difference between contexts. |
|
|
|
Number of continuous–categorical node pairs with an artificially introduced correlation difference between contexts. |
|
|
|
Number of continuous–continuous node pairs with both a mean shift and a correlation difference. |
|
|
|
Number of binary–binary node pairs with both a mean shift and a correlation difference. |
|
|
|
Number of categorical–categorical node pairs with both a mean shift and a correlation difference. |
|
|
|
Number of binary–continuous node pairs with both a mean shift and a correlation difference. |
|
|
|
Number of binary–categorical node pairs with both a mean shift and a correlation difference. |
|
|
|
Number of continuous–categorical node pairs with both a mean shift and a correlation difference. |
|
|
|
Magnitude of the mean shift introduced in shifted nodes. Must be ≥ 0. |
|
|
|
Target correlation difference introduced in correlated node pairs. Must be in [0, 1]. |
Real-World Data Parameters (real_world_data.*)
Only required when data_type = 'real'.
Parameter |
Type |
Default |
Description |
|---|---|---|---|
|
|
|
Absolute path to the CSV data file for context 1 (rows: samples, columns: variables). |
|
|
|
Absolute path to the CSV data file for context 2 (rows: samples, columns: variables). |
|
|
|
Absolute path to the metadata CSV file containing variable labels and data types. |
Differential Network Analysis Parameters (diff_net_analysis.*)
Parameter |
Type |
Default |
Description |
|---|---|---|---|
|
|
|
Path to the CSV samplesheet listing moDiNA configurations to run. Required when |
|
|
|
Node-level differential metric. Used when |
|
|
|
Edge-level differential metric. Used when |
|
|
|
Algorithm used to rank nodes and edges in the differential network. Used when |
|
|
|
Optional method to filter context-specific networks before differential analysis. One of: |
|
|
|
Parameter for the chosen filter method. Integer for |
|
|
|
Edge score used as the basis for filtering. One of: |
|
|
|
Rule applied when filtering edges. One of: |
|
|
|
Maximum path length used for the |
|
|
|
Type of statistical test used during context network inference. One of: |
|
|
|
Sentinel value in input data that represents missing / NA values. |
|
|
|
Multiple testing correction method applied to association p-values. One of: |
Usage
You can choose between the following profiles: conda, docker, singularity, slurm. The respective profile will be used to execute the pipeline with the appropriate containerization or job scheduling system. You can specify the profile using the -profile flag when running the Nextflow pipeline. For example, to run the pipeline using conda for package management and SLURM for distributed execution, you would use the following command:
Now, you can run the Nextflow pipeline using the following command:
nextflow run main.nf -profile conda,slurm
If you have stored the parameters in a params.yml file instead of the nextflow.config file, you can specify the path to this file using the -params-file flag:
nextflow run main.nf -profile conda,slurm -params-file path/to/your/params.yml
Pipeline Output
The results are stored in the specified output directory, which is organized by differential network analysis steps and evaluation steps.
The main output of the pipeline includes:
Simulated datasets for each simulation replicate (if
data_type = 'simulation').Inferred context-specific networks for each context and simulation.
Node and edge metrics files for each configuration and simulation.
Ranking results for each configuration and simulation.
Evaluation results.
In the following, we provide some examples of evaluation results generated by the Nextflow pipeline.
For evaluating the strength of the simulation, we can check the distribution of the introduced mean shifts and correlation differences across the simulated nodes and node pairs.
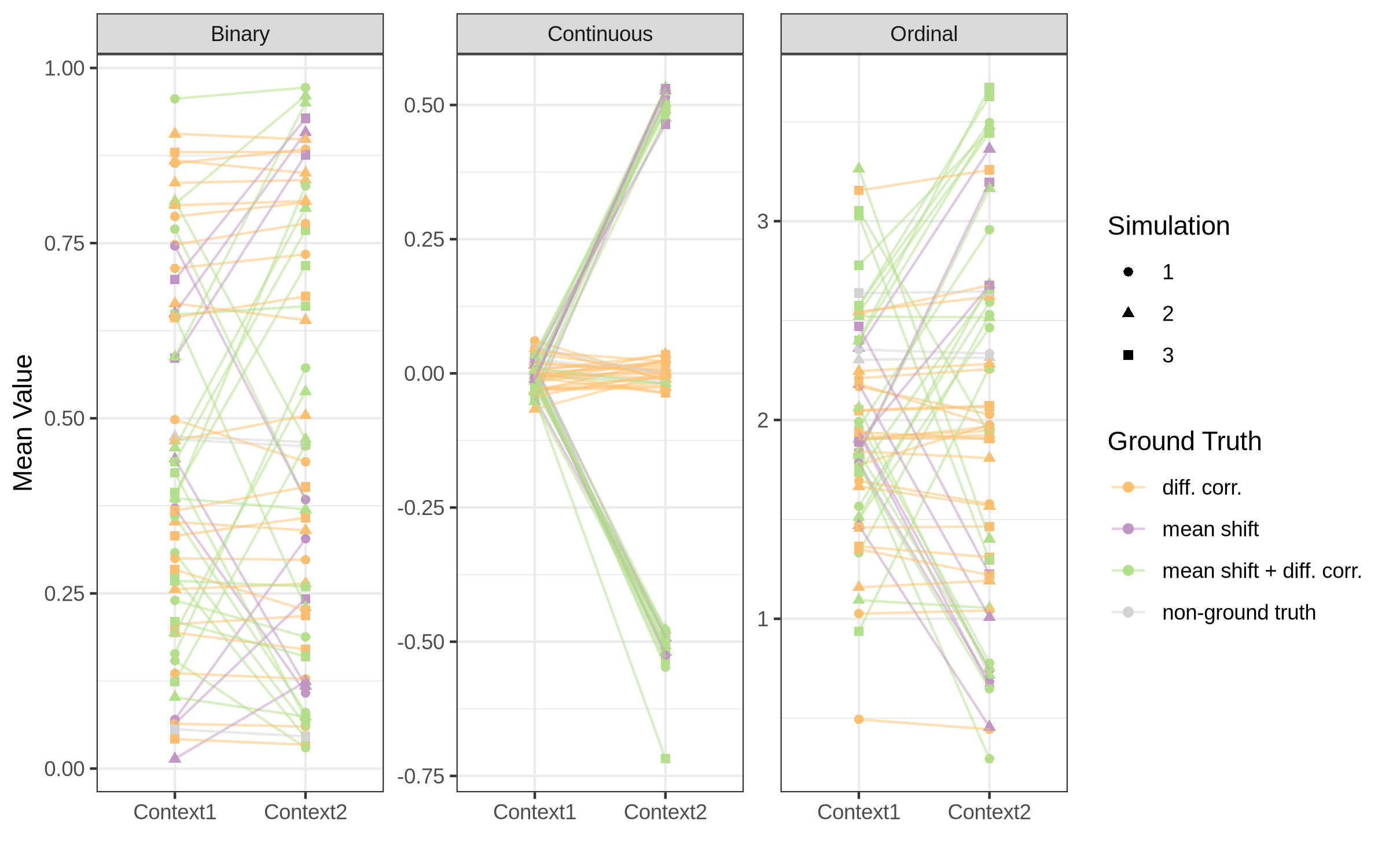
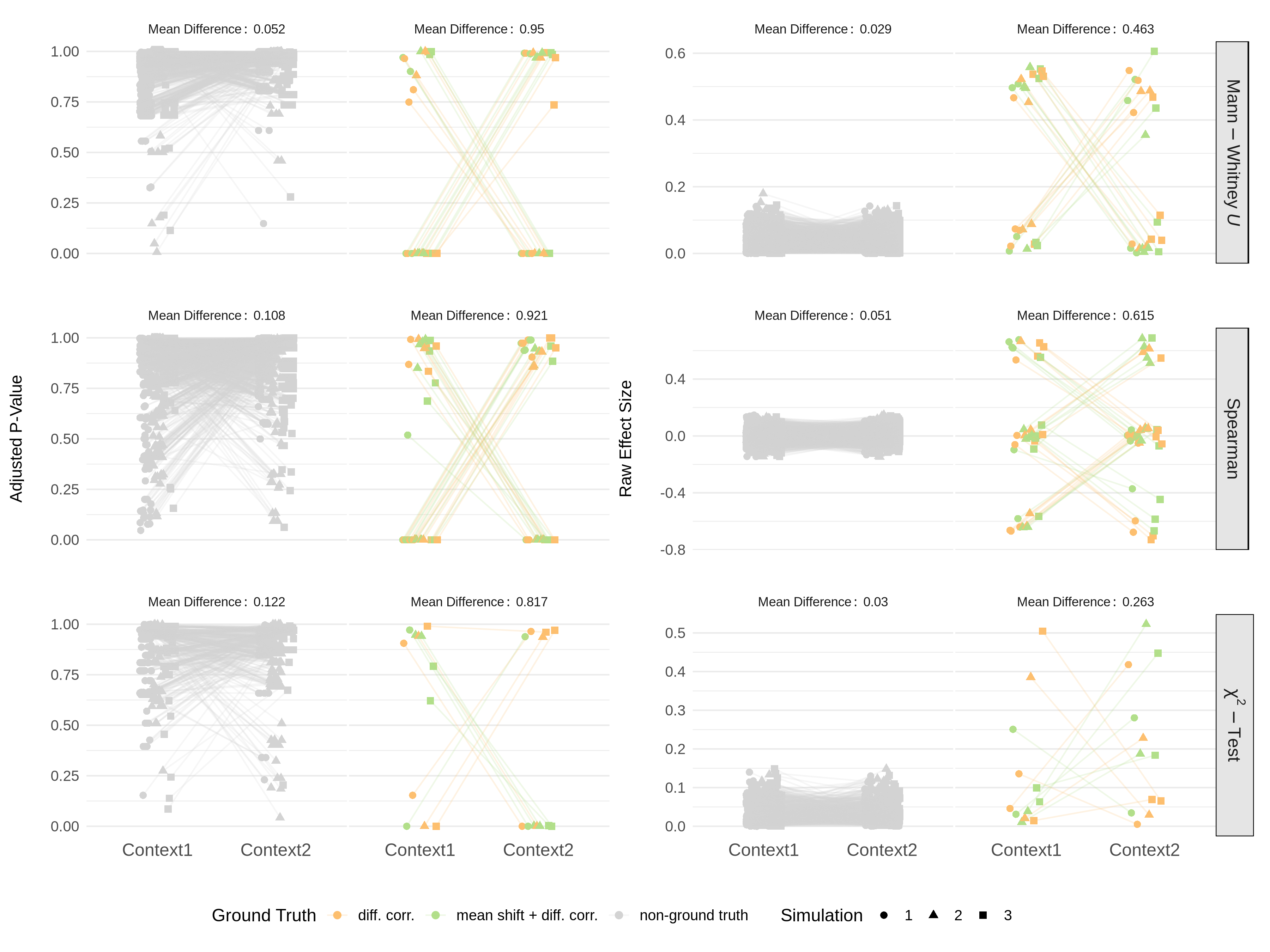
The performance of the different configurations can be evaluated using for example the AUC heatmap, which shows the AUC values for each configuration averaged across all simulation replicates.